Cattle as reservoirs of clinically relevant Enterococcus: A One Health genomic perspective
DOI:
https://doi.org/10.48797/sl.2026.401Keywords:
Selected Oral CommunicationAbstract
Background: Enterococcus spp. are opportunistic pathogens and One Health indicators of antimicrobial resistance (AMR) [1]. Cattle remain understudied reservoirs of multidrug-resistant (MDR) enterococci [2]. This study assessed the occurrence of clinically relevant antibiotic-resistant Enterococcus in cattle farms in Northern Portugal, comparing intensive dairy systems and native breeds under extensive conditions. Methods: A total of 120 fecal swabs were collected from Holstein-Friesian (n=60) and autochthonous breeds (n=60) across 20 farms (65 calves, 55 adults; 2023). Samples were pre-enriched with or without antibiotics (ampicillin/vancomycin/florfenicol) and plated on Slanetz–Bartley-agar, without/with those antibiotics. Identification (MALDI-TOF MS), susceptibility testing (9 antibiotics; EUCAST/CLSI) and whole-genome sequencing (Illumina) for 13 MDR isolates were performed. Prevalence was calculated per sample [3]. Results: Enterococcus were detected in 62% of samples, with frequent species co-occurrence, mainly E. faecium (Efm-67%), E. hirae (52%), E. faecalis (45%). Resistance to tetracycline (32%), erythromycin (30%), high-level-streptomycin (28%), ampicillin/ciprofloxacin (22% each), chloramphenicol (18%), linezolid (9%), and high-level-gentamicin (6%) was observed. MDR occurred in 26% of samples, mainly in calves and intensive farms. Linezolid resistance (LinR) genes (optrA, poxtA, cfr) were detected across species, mainly in intensive farms, with variable phenotypes (MIC 2-8mg/L). Ampicillin resistance was confined to E. faecium, including ST18 and ST80 lineages associated with hospital settings. One ST80 isolate carried vanHAX despite phenotypic susceptibility, suggesting a vancomycin-variable genotype. AmpR-ST80 CT9962 was restricted to one farm, whereas LinR-ST18-CT9964 spanned farms/cities (Fig.1). Efm showed mixed hospital/community virulence profiles, with clade A1 lineages (ST18/ST80/ST1693) harboring hospital-associated markers (ptsD/orf1481/IS16/full pili-clusters). Bacteriocin genes (0–6) were diverse; ST18 carried bacAS5 linked to hospital strains. Plasmid replicases (mostly Inc18) included hospital-associated plasmids in ST18 (rep_pNB2354p1) and ST80 (rep_pRUM). Conclusions: Cattle may act as reservoirs of enterococci resistant to critically important antibiotics. The detection of hospital-adapted E. faecium clones in intensive farming systems highlight potential transmission, supporting the urgent need for integrated One Health surveillance.
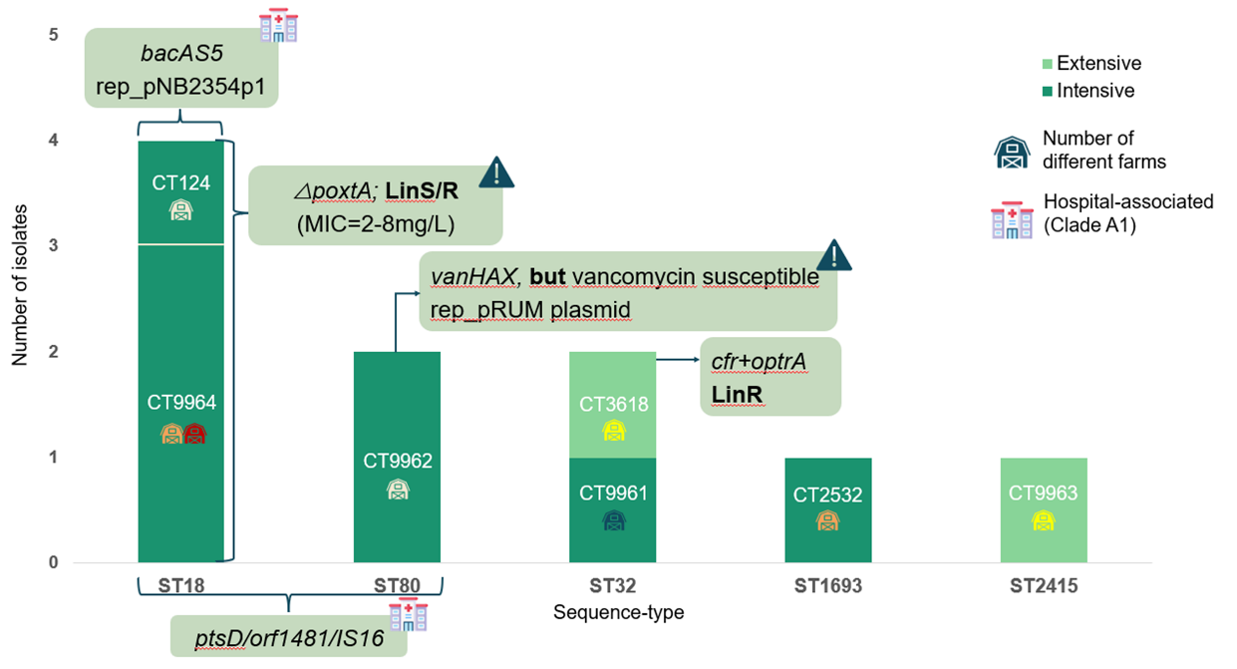
Figure 1. Distribution of E. faecium clones and hospital-associated markers across different farms based on whole-genome sequencing (WGS). Bars represent the number of isolates per sequence type (ST) and core-genome type (CT), colored by production system (intensive vs extensive). Icons indicate the number of farms where each clone was detected. Hospital-associated markers (e.g., ptsD, orf1481, IS16, pili clusters) were identified in specific lineages, particularly within clade A1. LinR= linezolid-resistant; LinS = linezolid-susceptible.
References
1. Torres, C. et al. Antimicrobial Resistance in Enterococcus spp. of animal origin. Microbiol Spectr 2018, 6(4), doi:10.1128/microbiolspec.arba-0032-2018
2. Gião, J. et al. Antimicrobial Susceptibility of Enterococcus Isolates from Cattle and Pigs in Portugal: Linezolid Resistance Genes optrA and poxtA. Antibiotics 2022, 11(5), 615, doi:10.3390/antibiotics11050615
3. Finisterra, L. et al. Industrial dog food is a vehicle of multidrug-resistant enterococci carrying virulence genes often linked to human infections. Int J Food Microbiol 2021, 358, 109284, doi:10.1016/j.ijfoodmicro.2021.109284
Downloads
Published
How to Cite
Issue
Section
License
Copyright (c) 2026 Inês M. Ribeiro, Ana C. Almeida-Santos, Maria J. Teixeira, A. Ribeiro, Carla Campos, Nuno V. Brito, Rui Dantas, Luís Pinho, Sandra Quinteira, Carla Miranda, Luísa Peixe, Carla Novais, Ana R. Freitas

This work is licensed under a Creative Commons Attribution 4.0 International License.
In Scientific Letters, articles are published under a CC-BY license (Creative Commons Attribution 4.0 International License), the most open license available. The users can share (copy and redistribute the material in any medium or format) and adapt (remix, transform, and build upon the material for any purpose, even commercially), as long as they give appropriate credit, provide a link to the license, and indicate if changes were made (read the full text of the license terms and conditions of use).
The author is the owner of the copyright.









